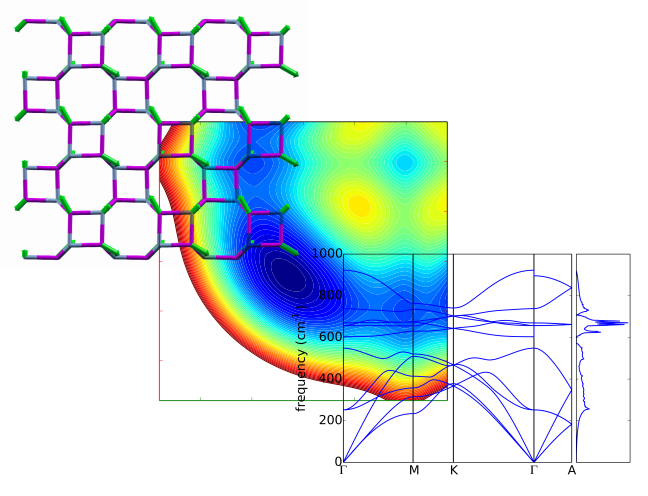
Features¶
Container classes for single unit cells (
Structure) and structure sequences such as molecular dynamics trajectories, relaxation runs or NEB paths (Trajectory). See Parsing code output and using containers.Classes to set up calculations (parameter studies) based on template input files for any kind of computational backend (
batch). See Parameter studies.Simple sqlite3 interface with convenience data extraction methods (
sql).Parsing of PWscf (QE), CPMD , CP2K and LAMMPS output into Python objects for easy access (
parse). See Parsing code output and using containers.Structure io: read cif, pdb, write axsf, cif, xyz (
io)Pythonic interface to external molecular viewers for interactive use: xcrysden, avogadro, jmol, VMD (
visualize).EOS fitting tools (
eos)Thermodynamic properties in the quasi-harmonic approximation from phonon density of states, QHA implementation (
thermo). See Quasi-harmonic approximation.MD analysis: radial pair distribution function (own implementation and VMD interface), RMS, RMSD (
crys).Velocity autocorrelation function and phonon DOS from MD trajectories (
pydos). See Velocity autocorrelation function and phonon DOS.Unit cell related tools: super cell building, coordinate transformation, k-grid tools, … (
crys).Functions and classes to extend numpy/scipy, e.g. N-dim polynomial fitting and a number of convenient 1D classes (polynomial, spline) with a common API (
num).N-dim radial basis function interpolation and fitting (
core,hyperopt). See Radial Basis Function interpolation an regression.Basic signal processing / fft related tools (
signal)Tools to handle matplotlib plots in scripts (
mpl)QE and LAMMPS calculators for ASE (
calculators)extensive test suite